Changelog
metapr2 3.0.0
Released: 2025-09-26
Database
New entries
- 8 new datasets
- 1650 new samples
Citations
- Sim et al. 2025. Temporal dynamics and biogeography of sympagic and planktonic photosynthetic microbial eukaryotes during the under-ice Arctic bloom. ISME Communications. ycaf075.
- Ribeiro et al. 2024. Arctic phytoplankton microdiversity across the marginal ice zone: Subspecies vulnerability to sea-ice loss. Elementa: Science of the Anthropocene. 12:00109.
- Egge et al. 2021. An 18S V4 rRNA metabarcoding dataset of protist diversity in the Atlantic inflow to the Arctic Ocean, through the year and down to 1000 m depth. Earth System Science Data. 13:4913–28.
- Hörstmann et al. 2024. Biogeographic gradients of picoplankton diversity indicate increasing dominance of prokaryotes in warmer Arctic fjords. Commun Biol. 7:1–11.
- Šupraha et al.. 2022. Diversity and biogeography of planktonic diatoms in Svalbard fjords: the role of dispersal and Arctic endemism in phytoplankton community structuring. Elementa: Science of the Anthropocene. 10:00117.
- Ong et al. (2025), Consistent cell-specific carbon fixation rates by small eukaryotic phytoplankton in contrasting nutrient-limited conditions. Limnol Oceanogr, 70: 162-177. https://doi.org/10.1002/lno.12751
- Décima et al. 2023. Salp blooms drive strong increases in passive carbon export in the Southern Ocean. Nat Commun. 14:1–16.
metapr2 2.1.0
Released: 2023-05-16
metapr2 2.0.0
Released: 2022-11-23
Database
59 datasets
18 new datasets
Polar
- Arctic - 2012 (Kilias 2020)
- Amundsen_Sea ASPIRE cruise - 2010-2011
- Fram Strait - 2014
- Palmer Station Antarctic - LTER - 2014
- Arctic and Scotian Shelf - 2009-2011
- Baffin Bay - 2008-2018
- Southern Ocean - 2017
- Fram observatory 2016
- Antarctic Peninsula - 2012-2016
Coastal
- Roscoff Astan - 2009-2011
- Roscoff Astan - 2012-2016
- SE Asia Tsunami deposits
- Baltic Sea Gdansk Gulf - 2012 Hapto
- Coral Infecting Apicomplexan
- Zostera marina - British Columbia - 2015
- Coral Singapore - 2018
Clustering
An option is now provided to use either all ASVs or clustered ASVs on the welcome screen. ASVs are clustered at 100% identity with VSEARCH –id 1.00 See the metaPR2 paper for more information.
It is also possible to use version 1.0 of the database by entering v1 on the welcome screen.
Web application
New Panel: Taxonomy
This new panel provides a table with all the taxa present in the current metaPR2 version with the number of ASV for each species. The table can be easily searched.
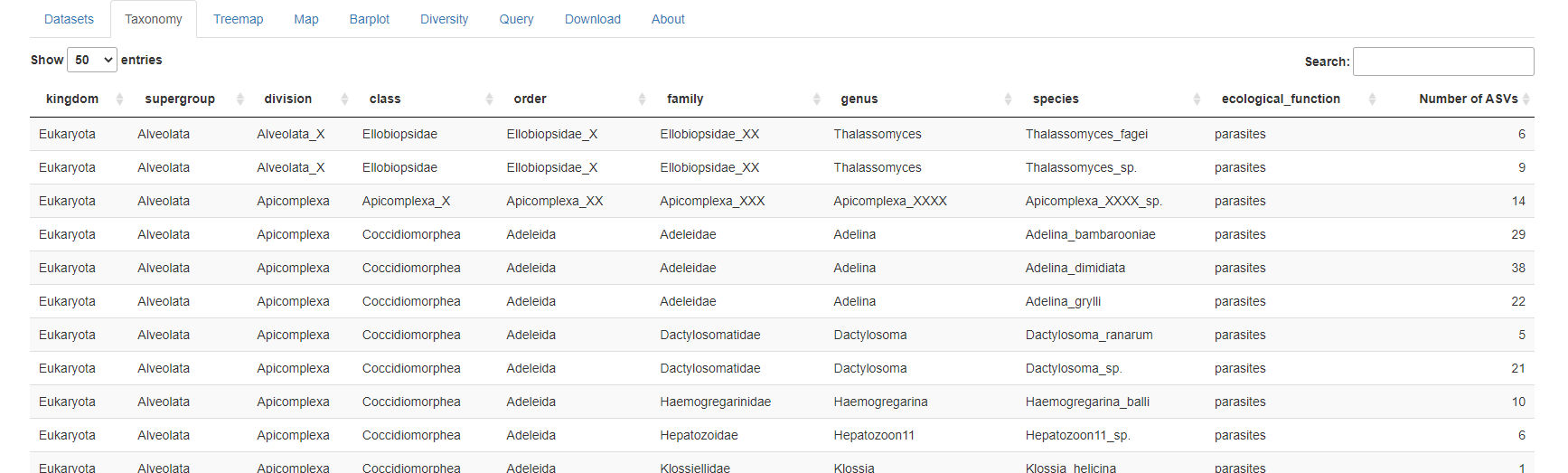
Minor changes
Information about database is provided on left panel
New button to disconnect the application and reloading.
Maximum number of samples for Phyloseq: 2000.
Taxonomy is constructed from all the samples and not only samples selected.
Option to use clustered ASVs in Welcome panel.
Barplots. The right side of the graph indicates, for each parameter range, the number of samples that fall into that range as well as the number of samples that contain the taxa selected.
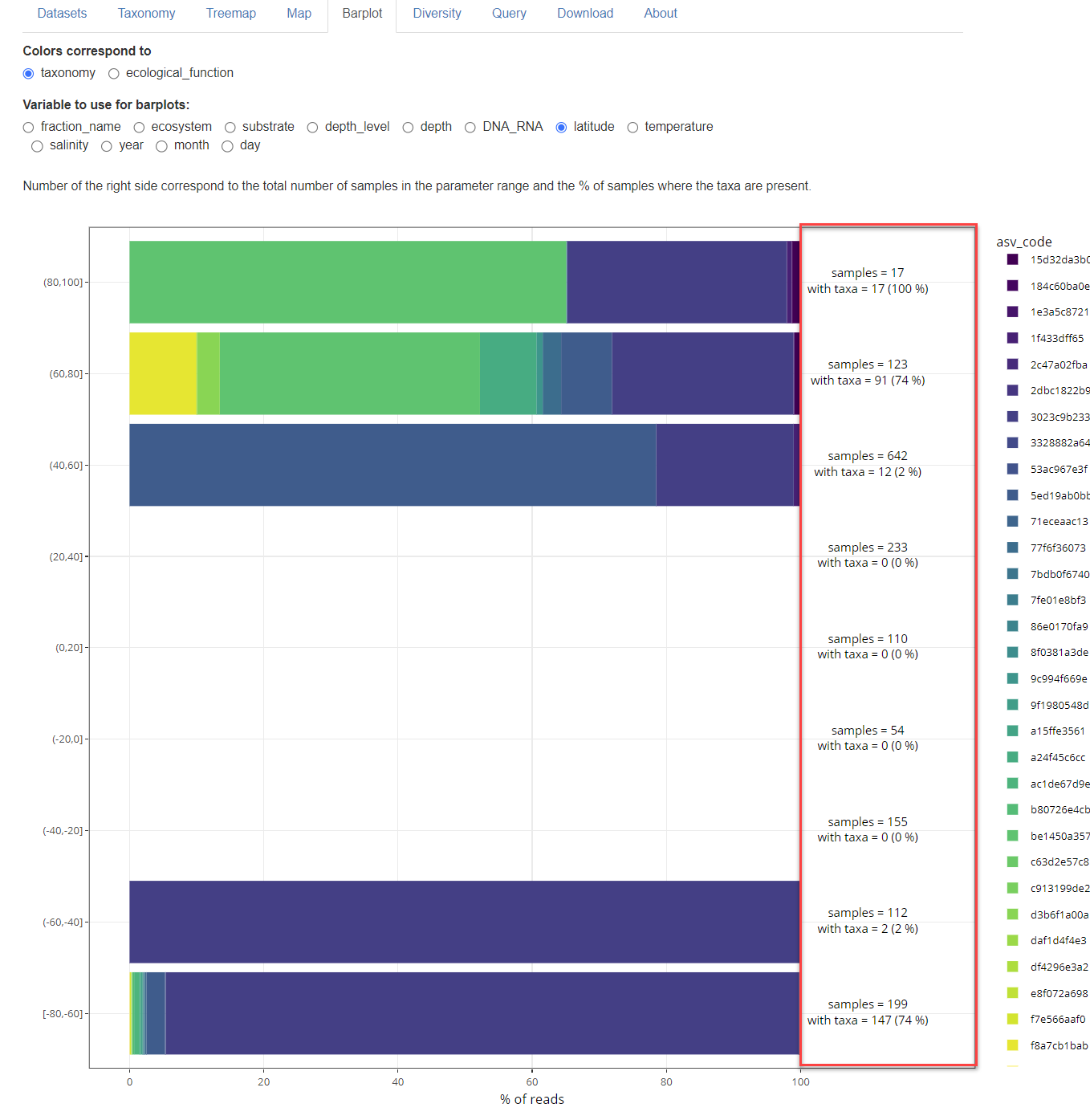
metapr2 1.0.2
Released: 2021-12-14
metapr2 1.0.1
Released: 2021-11-22
Tabs of application
Documentation
- Using pkgdown: https://pr2database.github.io/metapr2-shiny/